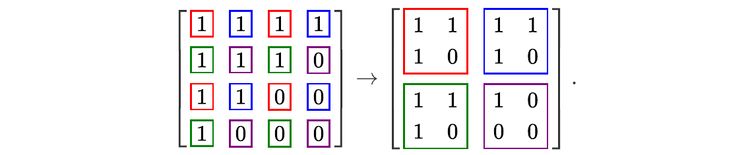
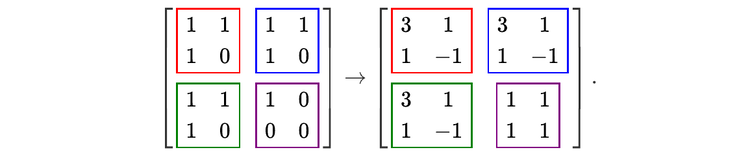
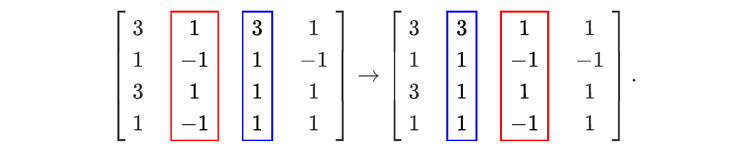
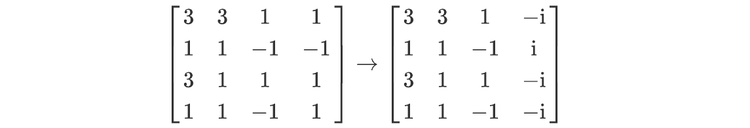
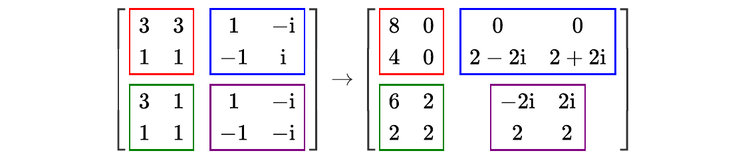
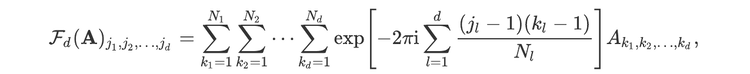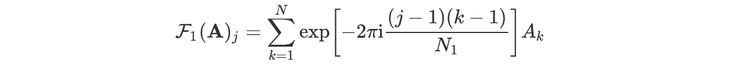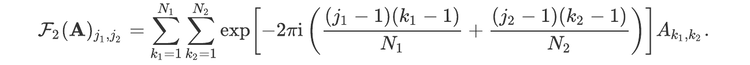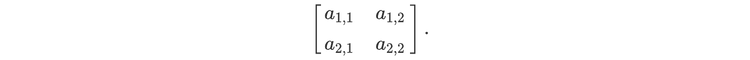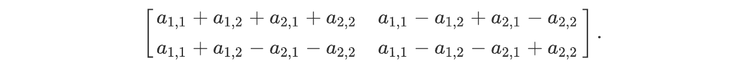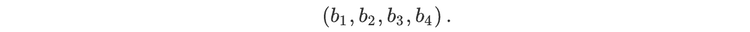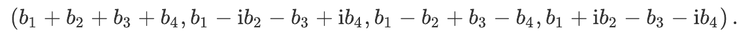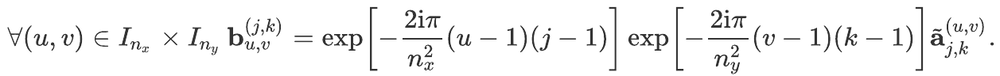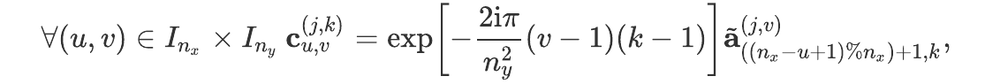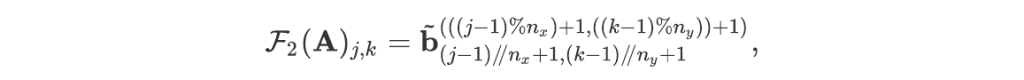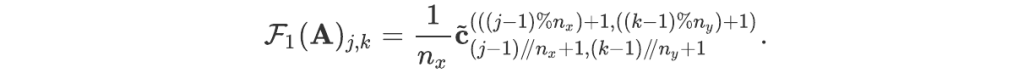For readers who may be more familiar with code than with mathematical notations, here is a Python function implementing this algorithm. It takes as argument a NumPy array, img, with at least two dimensions, and returns the FT along the last dimension. In this script,
-
nx and ny give the shape of the input for the device (for instance, in the above example, they would both be equal to 2),
-
device_FT is the 2D Fourier transform done by the optical device (if you want to try this script, you may use np.fft.fft2, as shown in the examples below),
-
the last two dimensions of the input must have lengths equal to the squares of nx and ny, respectively.
import numpy as np
def FFT1D(img):
'''
1D Fourier transforms
Arguments:
img: 2+ dimensional real or complex array of shape (..., nx**2, ny**2)
'''
try:
assert (img.shape[-1]) == nx**2
assert (img.shape[-2]) == ny**2
except:
print('Wrong image shape: expected ({},{}); got {}'
.format(nx**2, ny**2, img.shape[-2:]))
return
# shape of the batch
n_imgs = img.shape[:-2]
d_imgs = tuple(range(len(n_imgs)))
# create the Fourier transforms of the sub-arrays
a = img.reshape(n_imgs + (nx,nx,ny,ny)).transpose(d_imgs + (-3,-1,-4,-2))
tilde_a = device_FT(a)
# multiply by the complex numbers with modulus 1
j_jp = np.arange(ny)[:,np.newaxis].dot(np.arange(ny)[np.newaxis,:])
ey = np.exp(-2.*np.pi*1j/(ny*ny) * j_jp)
tilde_a_p = tilde_a * ey[np.newaxis,:,np.newaxis,:]
# some rearrangement of the lines
tilde_a_p[...,1:,:] = tilde_a_p[...,-1:0:-1,:]
# transpose
bar_a = tilde_a_p.transpose(d_imgs + (-4,-1,-2,-3))
# second round of Fourier transforms
tilde_bar_a = device_FT(bar_a)
# reconstruct the result
tilde_A = tilde_bar_a.transpose(d_imgs + (-2,-4,-1,-3))\
.reshape(n_imgs + (nx*nx,ny*ny)) / ny
return tilde_A
Here is an example of use in an IPython instance (after copying the above code in the system clipboard; notice that Python uses the letter j instead of i for the square root of -1):
In [1]: %paste
...
In [2]: device_FT = np.fft.fft2; # use the NumPy fft2 function
In [3]: nx=2; ny=2; # set nx and ny
In [4]: a = np.array([[1,1,1,1],
...: [1,1,1,0],
...: [1,1,0,0],
...: [1,0,0,0]])
In [5]: print(FFT1D(a).round(15))
[[ 4.+0.j 0.+0.j 0.+0.j 0.+0.j]
[ 3.+0.j -0.-1.j 1.+0.j 0.+1.j]
[ 2.+0.j 1.-1.j 0.+0.j 1.+1.j]
[ 1.+0.j 1.+0.j 1.+0.j 1.+0.j]]
Each line of the output is the discrete FT of the corresponding line of the array a. This is, in itself, not very impressive: the function np.fft.fftwould have done exactly the same thing! What is more interesting is that, to perform this calculation, the function FFT1D has only done small 2-dimensional FTs of size 2 by 2—exactly what a (very small) optical device could do!
It is easy to verify that it also works for larger arrays. For instance, changing the values of nx and ny to 3, we can compute the FTs of the arrays (1,1,1,1,1,1,1,1,1), (2,2,2,2,2,2,2,2,2), …, (9,9,9,9,9,9,9,9,9):
In [6]: nx=3; ny=3; # changing the values of nx and ny
In [7]: a = np.ones((9,9)) * np.arange(1,10)[:,np.newaxis]; print(a) # input
[[1., 1., 1., 1., 1., 1., 1., 1., 1.],
[2., 2., 2., 2., 2., 2., 2., 2., 2.],
[3., 3., 3., 3., 3., 3., 3., 3., 3.],
[4., 4., 4., 4., 4., 4., 4., 4., 4.],
[5., 5., 5., 5., 5., 5., 5., 5., 5.],
[6., 6., 6., 6., 6., 6., 6., 6., 6.],
[7., 7., 7., 7., 7., 7., 7., 7., 7.],
[8., 8., 8., 8., 8., 8., 8., 8., 8.],
[9., 9., 9., 9., 9., 9., 9., 9., 9.]]
In [8]: print(FFT1D(a))
[[ 9.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j]
[18.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j]
[27.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j]
[36.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j]
[45.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j]
[54.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j]
[63.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j]
[72.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j]
[81.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j 0.+0.j]]
You can check that the result is correct (the FT of a uniform array has just one non-vanishing coefficient, which is the sum of the input coefficients).
Finally, let us do a check with a 100 by 100 random matrix. It would, of course, be quite cumbersome to check the result by hand. Instead, let us compare the result with that of the function np.fft.fft, which is the NumPy one-dimensional FT implementation, and compute the mean absolute value of the difference:
In [9]: nx=10; ny=10;
In [10]: a = np.random.uniform(size=(100,100))
In [11]: print(np.abs(FFT1D(a) - np.fft.fft(a)).mean())
1.1224604726207545e-15
The difference is not exactly 0; this is because the numbers Python work with do not have an infinite precision. What this result shows is that the two ways of computing the FTs give the same result up to the numerical accuracy of the engine. (You will probably get a slightly different number if you try this code. This is because the matrix a is chosen randomly each time the code is run. But the result should be smaller than 10⁻¹⁴.)